SEARCH
Search
Indole test
-
General
The indole test is a biochemical test performed on bacterial species to determine the ability of the organism to convert tryptophan into indole. This division is performed by a chain of a number of different intracellular enzymes, a system generally referred to as "tryptophanase."
Biochemistry
Indole is generated by reductive deamination from tryptophan via the intermediate molecule indolepyruvic acid. Tryptophanase catalyzes the deamination reaction, during which the amine (-NH2) group of the tryptophan molecule is removed. Final products of the reaction are indole, pyruvic acid, ammonium (NH4+) and energy. Pyridoxal phosphate is required as a coenzyme.
Performing a Test
Two methods are in use;
1) a conventional tube method requiring overnight incubation, which identifies weak indole producing organisms and
2) a spot indole test, which detects rapid indole producing organisms
Procedure of Conventional Tube method for Indole Test
a. Inoculate the tryptophan broth with broth culture or emulsify isolated colony of the test organism in tryptophan broth.
b. Incubate at 37°C for 24-28 hours in ambient air.
c. Add 0.5 ml of Kovac’s reagent to the broth culture.
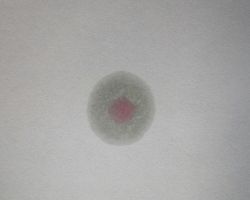
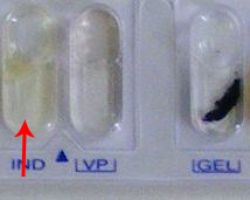
Expected results:
♦ Positive: Pink colored rink after addition of appropriate reagent
♦ Negative: No color change even after the addition of appropriate reagent. e.g. Klebsiella pneumoniae
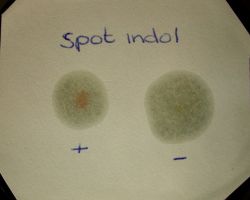
Spot Indole Test
It is used to determine the presence of the enzyme tryptophanase. Tryptophanase breaks down tryptophan to release indole, which when reacts with cinnamaldehyde produces a blue-green compound. The absence of enzyme results in no color production (i.e. indole negative).
Procedure of Spot Indole Test
♦ Saturate a piece of filter paper with the 1% paradimethylaminocinnamaldehyde reagent.
♦ Use a wooden stick or bacteriologic loop to remove a small portion of a bacterial colony from the agar surface and rub the sample on the filter paper.
Note: The bacterial inoculum should not be taken from MacConkey agar because the color of lactose fermenting colonies on this medium can interfere with test interpretation.
Result:
1) Positive: Development of a blue color within 30 seconds. Most indole positive organism turn blue within 30 seconds.
2)Negative: No color development or slightly pink color.
Uses:
Spot indole test along with Gram stain result and colony characteristics can assist in the rapid identification of isolates. For example
1) A flat, dry lactose fermenting (pink) colony on MacConkey agar that also spot indole positive and oxidase negative can be reported presumptively as E.coli.
2) Organisms that swarm on 5% sheep blood agar, exhibit a characteristics odour, and are oxidase negative can be presumptively identified as Proteus spp. With further testing by spot indole, the positive isolates may be presumptively reported as Proteus vulgaris and the negative ones as Proteus mirabilis.
Indole positive organisms: Most strains of E.coli, P. vulgaris, M. morganii and Providenica are indole positive.
Point to remember: Indole test can also aid in species differentiation.
1) Klebsiella species: Klebsiella oxytoca is indole positive whereas Klebsiella pneumoniae is indole negative.
2) Citrobacter species: Citrobacter Koseri is indole positive where as Citrobacter freundii is indole negative
3) Proteus species: Proteus Vulgaris is indole positive whereas Proteus mirabilis is indole negative
I used to remember above mentioned information with the help of MNEOMONIC, which can be useful for you.
For this remember the phrase “OK VIP”
Where: O means: Oxytica (Klebsiella oxytoca), K means (koseri-i.e. Citrobacter koseri), V means vulgaris (Proteus vulgaris) and IP means Indole Positive.
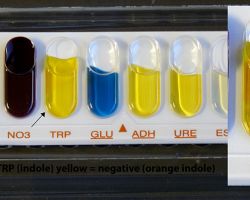
Delftia acidovorans
Identification of the microorganism can be performed using a simple orange indole reaction test. With the addition of Kovac’s reagent, the organism produces anthranilic acid using tryptophan.
This results in a pumpkin-yellow/orange colour in the media, which is characteristic for D acidovorans
-
History
-
Related
-
References
Wikipedia
https://microbeonline.com/indole-test-principle-procedure-results/
Photo
Wikipedia
MMIZ, ErasmusMC, Rotterdam
https://microbiologyinfo.com/indole-test-principle-reagents-procedure-result-interpretation-and-limitations/

-250x200.jpg)
-250x200.png)